Submitting a gene list for Literature Lab™ PLUS analysis.
A Primer on using the Literature Lab™ Gene List Editor.
These instructions use a small spreadsheet with a 17 gene molecular signature. The journal article reference for the gene list is provided on the spreadsheet. A small gene list is used in order to facilitate a quick exercise.
Download the file to your computer. One gene symbol error has been made in the file so that we can introduce basic Literature Lab™ Gene Thesaurus tools for validating gene data sets - 'RUNX1' has been changed to 'RUNX'.
We will use this file twice:
First, to illustrate a how to import a spreadsheet with a list of genes and submit it for Literature Lab™ PLUS analysis..
Second, to illustrate importing a gene set with up- and down-regulated genes and the process recommended by Acumenta Biotech for using Literature Lab™ PLUS to perform functional analysis on this type of data.
Literature Lab™ imports a variety of file types and input methods:
.xls, .xlsx, text files and comma/tab delimited files.
Gene lists can also be entered by typing, and genes can be added to an imported list by typing.
Queries to the Gene Thesaurus can be used to find gene symbols and enter them on a list.
Multiple gene lists can be imported and combined into a single file for Literature Lab™ PLUS analysis.
Example 1: Import a gene list, validate it, and submit it for Literature Lab™ PLUS analysis.
Start the Gene List Editor. Make sure the Data Set Directory setting has been verified by the test done previously, which you can find here if you haven't already done this..
Click the button on the lower right: 'Create a New Gene List' to get started.
Click the button on upper right: 'Import a Gene List’. An informational window will appear, click 'Yes'.
The spreadsheet has human gene symbols, the default on this screen. Other annotations are supported and you can explore them in the drop down menu at the field: 'Original ID Type:'.
Use the file browser to specify the location of the spreadsheet on your computer and click 'Import'.
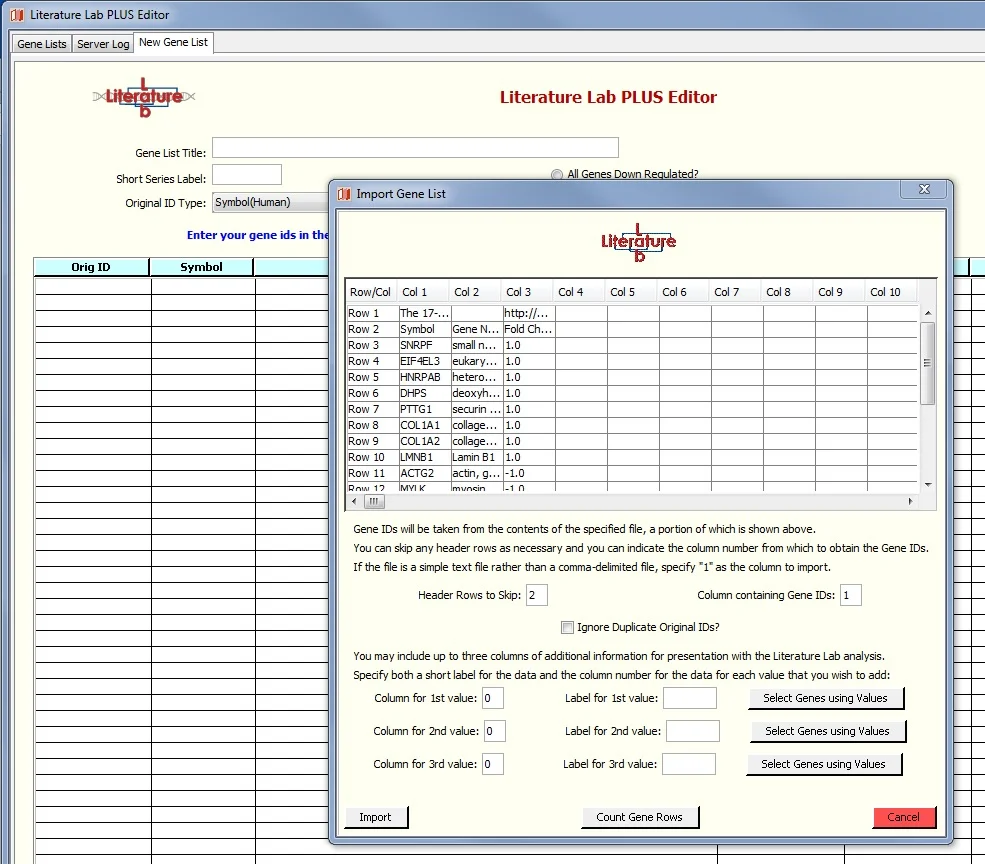
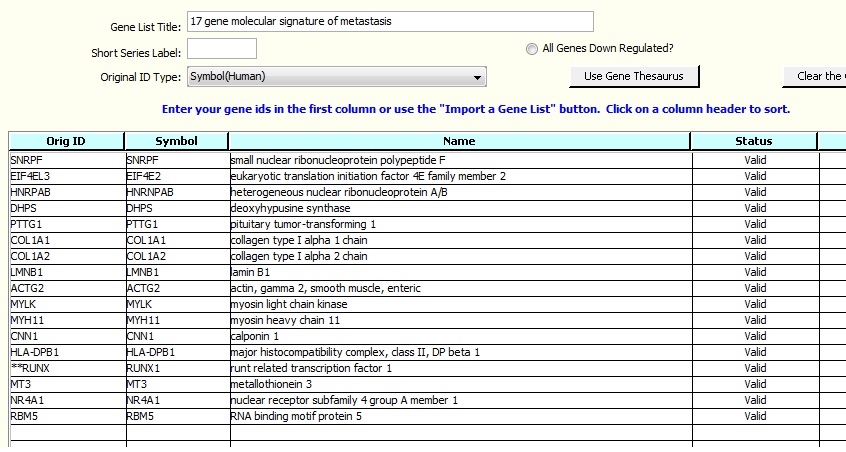
You will be presented with a window into the spreadsheet per the screen image below. Set the value in 'Header Rows to Skip:' to 2 because the first two rows do not contain genes. The gene symbols are in column 1. The Fold Change Column is not used in this exercise.
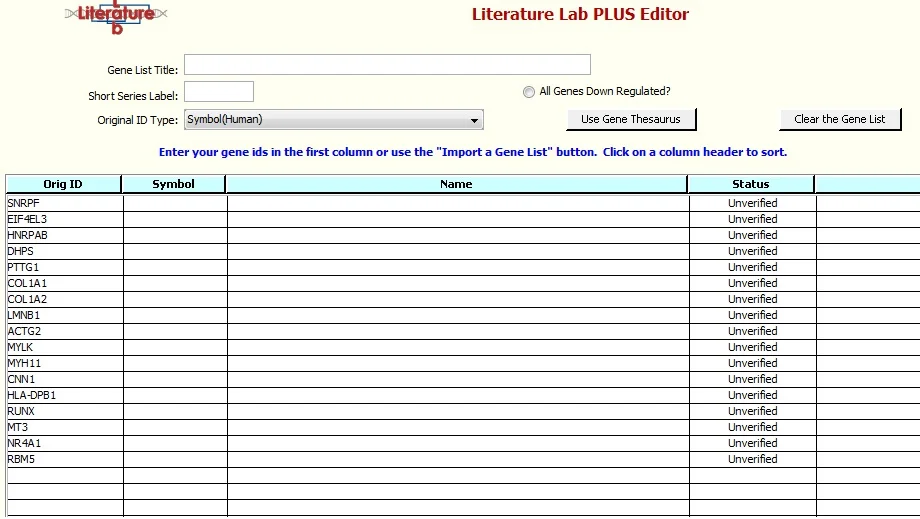
7. Click 'Import', see this screen:
Click 'Validate the Gene List'. The Gene Thesaurus will load. When asked 'If a gene symbol is ambiguous, do you want to automatically select the gene for which the symbol is the primary gene symbol?' select 'Yes'.
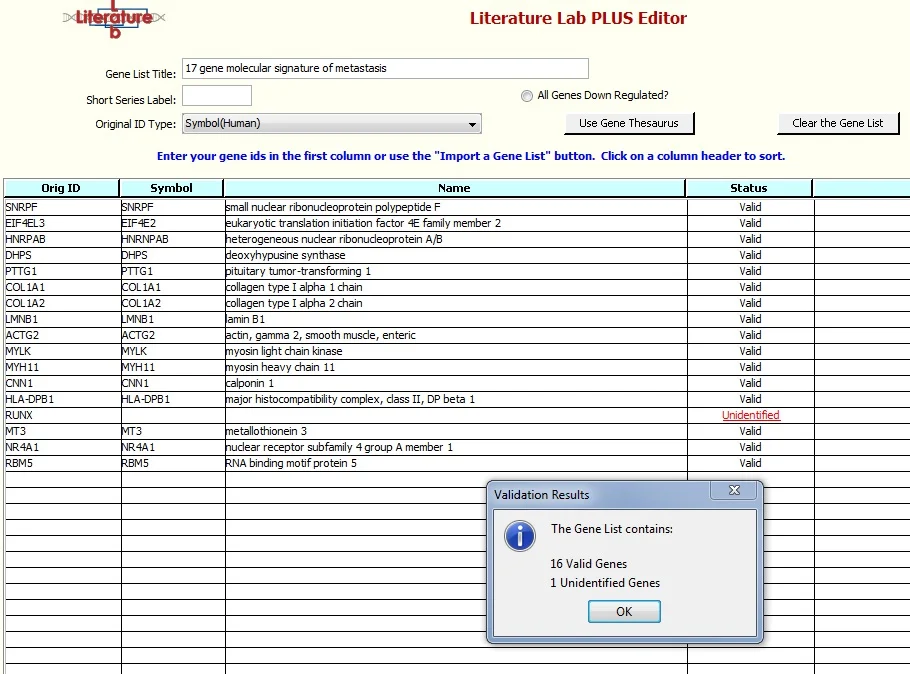
The gene symbols will be checked against the Gene Thesaurus and the gene names will be entered. 'RUNX', wasn't found in the Gene Thesaurus and the Status for this entry is 'Unidentified'.
Click "'OK' and click on 'Unidentified', this is a link into the Gene Thesaurus.
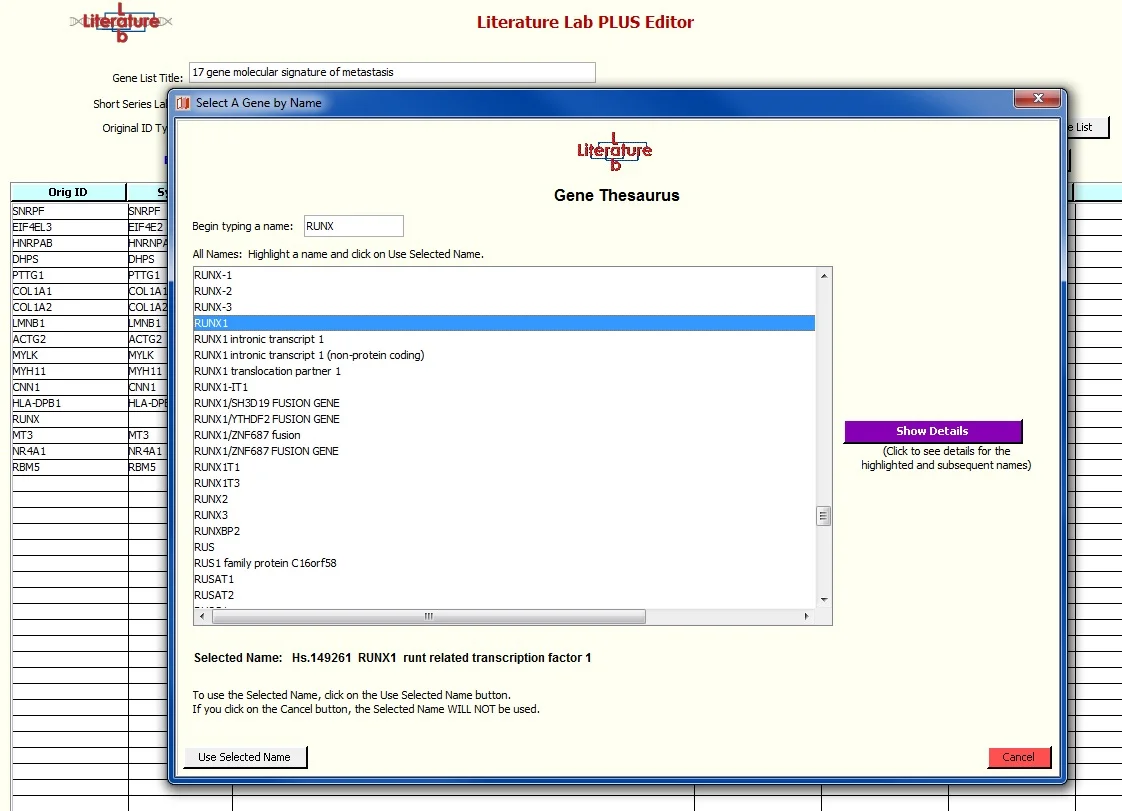
The Gene Thesaurus window opens to a location close to the unidentified entry. Use the up/down cursor keys or click to select the correct gene. If the gene names are in a column on your import spreadsheet you can use this to help you find the correct gene if you are unsure which symbol to select. Click on ‘Show Details’ to see the name and aliases for the gene that is currently highlighted.
After highlighting 'RUNX1' click 'Use Selected Name'.
RUNX1 has been entered into the Symbol column, the name has been fetched and the Status is set to Valid. The two asterisks on RUNX indicate that the value from the imported spreadsheet has been modified.
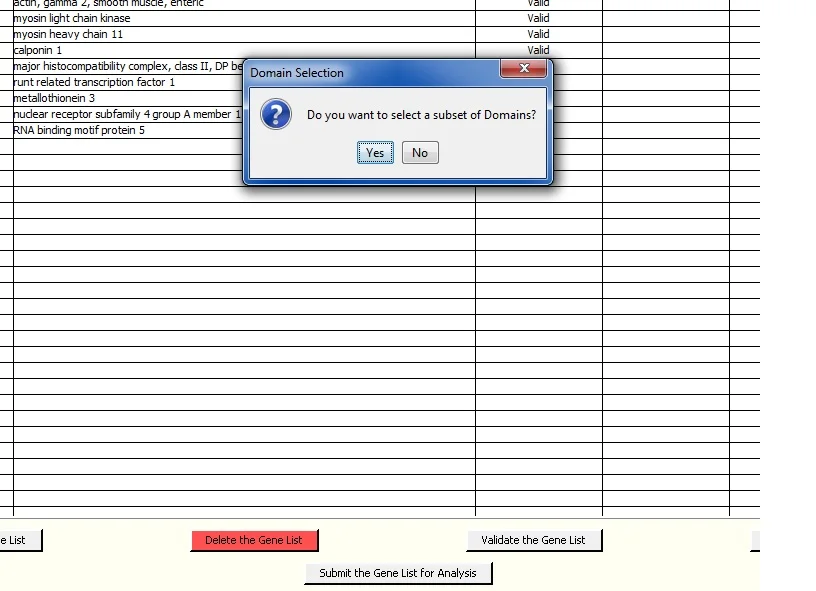
Click 'Submit the Gene List for Analysis' at the bottom center of the screen to put the gene list into a queue for analysis. The following query window is presented - you can run the analysis for all Domains if you click 'No'. Click 'Yes' to select a subset of Domains. The 'Yes' screen gives you the option to select a default subset of Domains. This is useful if you regularly want to use the same subset of domains for the analysis..
The Editor will present a status message and switch to the analyzer Process Log screen. Go back to the Gene Lists tab if you want to submit another gene list for analysis. You can continue to submit gene lists for analysis, the Literature Lab™ PLUS Analyzer processes gene lists in the order submitted. If you are done using the Editor, close it.
Start the Analyzer if it isn't already running. Click on the Process Log tab. You will see updates flowing into the log. There are 6 statuses for each Domain (12 log entries per Domain - Started/Completed for each status point), and the log will move right along on a fast computer. Some of the Domains have thousands of terms (Diseases, Chemicals & Drugs, Substances), so don't think it is frozen when it works on these Domains.
You can open Literature Lab™ PLUS and you will see the new experiment on the list that comprises the opening screen. The Run Date field will be blank until the analysis is completed
Nota Bene: Submitting a gene list for analysis does not in itself start the Literature Lab™ PLUS statistical analysis on the gene list. You must start the LitLab+ Analyzer process via the startup icon. This program can be running while you enter additional gene lists for analysis - all Literature Lab™ programs can run concurrently.
example 2: Importing, validating and submitting gene lists consisting of two differentiated subsets, e.g.: up- and down-regulated genes.
The Literature Lab™ Gene List Editor has features that simplify importing spreadsheets with supplemental data values falling on either side of zero (or any specified value). Acumenta Biotech recommends that all data sets with +/- data, e.g.: fold change data, be run as two separate gene sets and that Literature Lab™ PLUS viewer comparison features be used to identify functional similarities and differences among the two subsets. Instructions for using these features are provided in the PLUS Viewer Help files. Acumenta Biotech does not recommend running only a single "blended" analysis on the entire gene list.
The Gene List Editor can be used to specify ranges to be imported for analysis. For this exercise we will show how to import a single spreadsheet with up- and down-regulated data and Submit separate Literature Lab™ PLUS analyses for each subset.
The spreadsheet downloaded for this exercise contains a Fold Change column which we ignored in the first example. Values in this column have been set to either +1 or -1 in lieu of the original fold change data that was not provided in the table in the article.
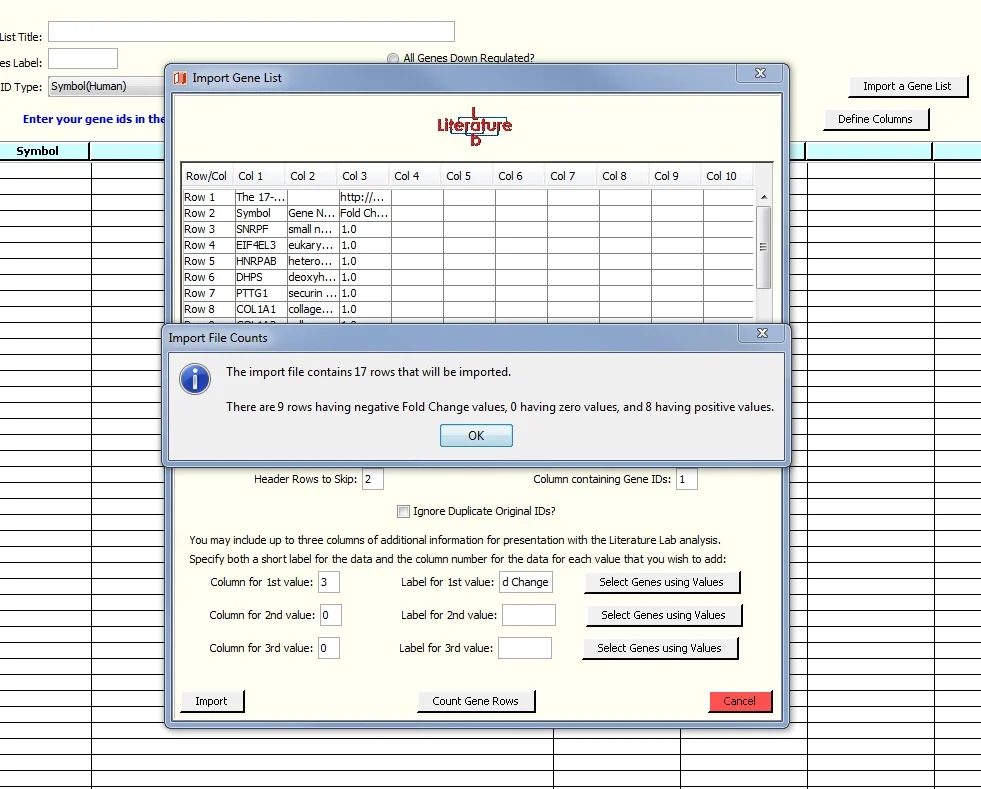
Import the spreadsheet as shown above, but add the Fold Change data to the Import window. Note that as before we have entered '2' in the 'Header Rows to Skip' field and '1' in the Gene ID Column. As shown below, we have added information about column 3 of the spreadsheet.
Finally we clicked on the 'Count Gene Rows' button to open a window providing a tally of genes having negative, positive or zero Fold Change values.
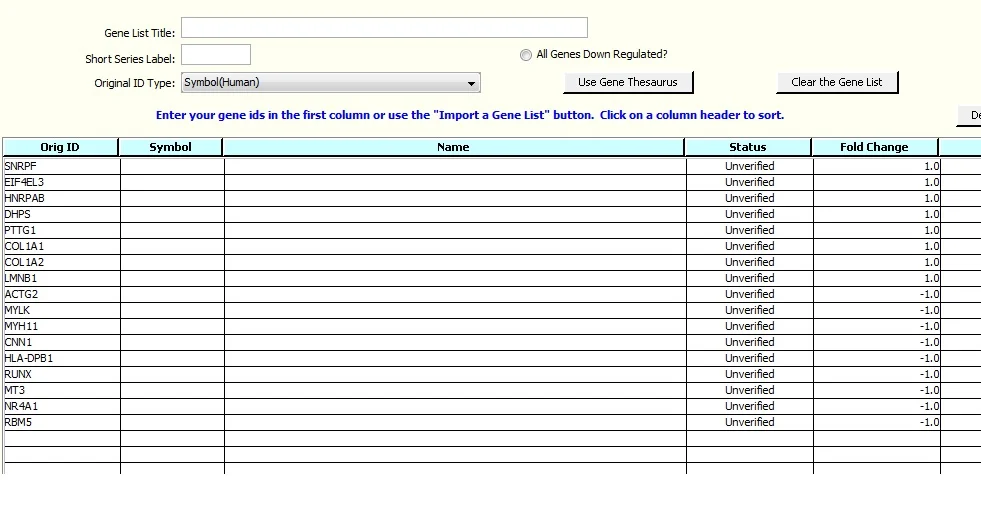
Click on 'Import', and note that the Editor has the fold change data. Up to three columns of supplemental data can be imported. Ranges can be specified using the 'Select Genes Using Values' button, e.g.: if you want to exclude genes with fold change values in a specified range. See Example 3 for an example of removing genes with selected fold change values.
Users frequently import fold change and p-value data. This data is not used in the Literature Lab™ analysis. It is useful for adding context in the PLUS Viewer, and results can be ranked by the these values. See the the Help files in Literature Lab™ PLUS for more information.
Validate the gene list and repair the error in the imported symbol for RUNX1 as described in Example 1, above.
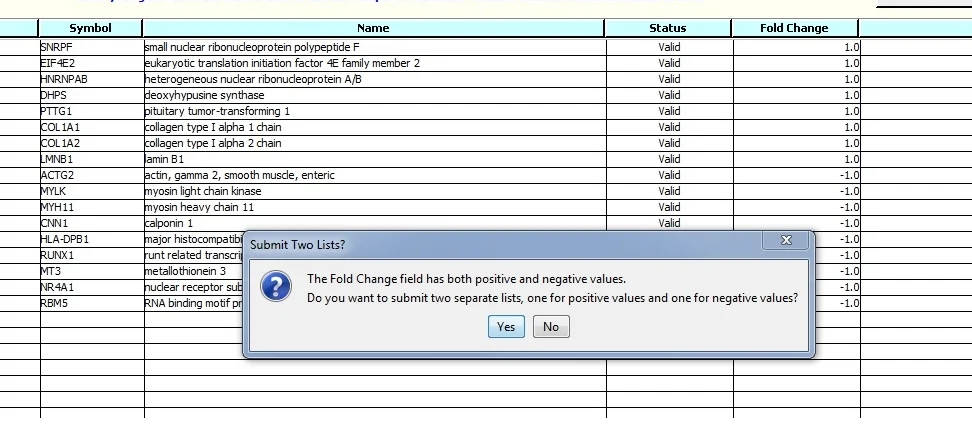
When we Submit the data for Analysis we see a message about the values in the Fold Change field.
Click "Yes". Two gene lists will be submitted for analysis. '_Down' and '_Up' are appended to the original file name.
Proceed to the next page for instructions about specifying fold change cutoffs.