Submitting a gene list for Literature Lab™ PLUS analysis.
Example 3: Using the Gene List Editor to specify fold change cutoffs.
The Literature Lab™ Gene List Editor has features that simplify importing spreadsheets with supplemental data values such as fold change, p-values, etc.. Acumenta Biotech recommends that all data sets with +/- values, e.g., fold change data, be run as two separate gene sets, and that the Literature Lab™ PLUS viewer comparison features be used to identify functional similarities and differences among the two subsets. Instructions for using these features are provided in the PLUS Viewer Help files. We do not recommend running only a single 'blended' analysis on the entire gene list.
In this illustration, we show the Literature Lab Gene List Editor functions for validating and submitting data subsets differentiated by, in this example, positive and negative fold change.
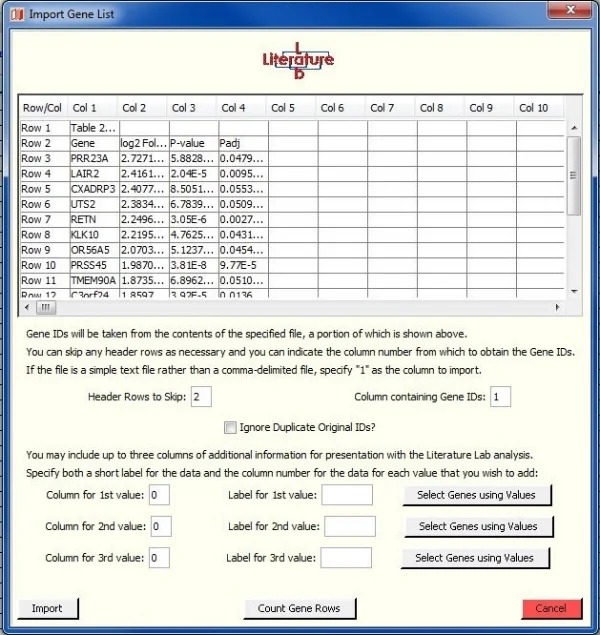
The import preview window
The Gene List Editor can import up to 3 columns of additional data. In this exercise, we will import the fold change data.
The preview window shows that column 2 of the import spreadsheet has fold change data.
The genes are in column 1. Note that there are two header rows in the spreadsheet.
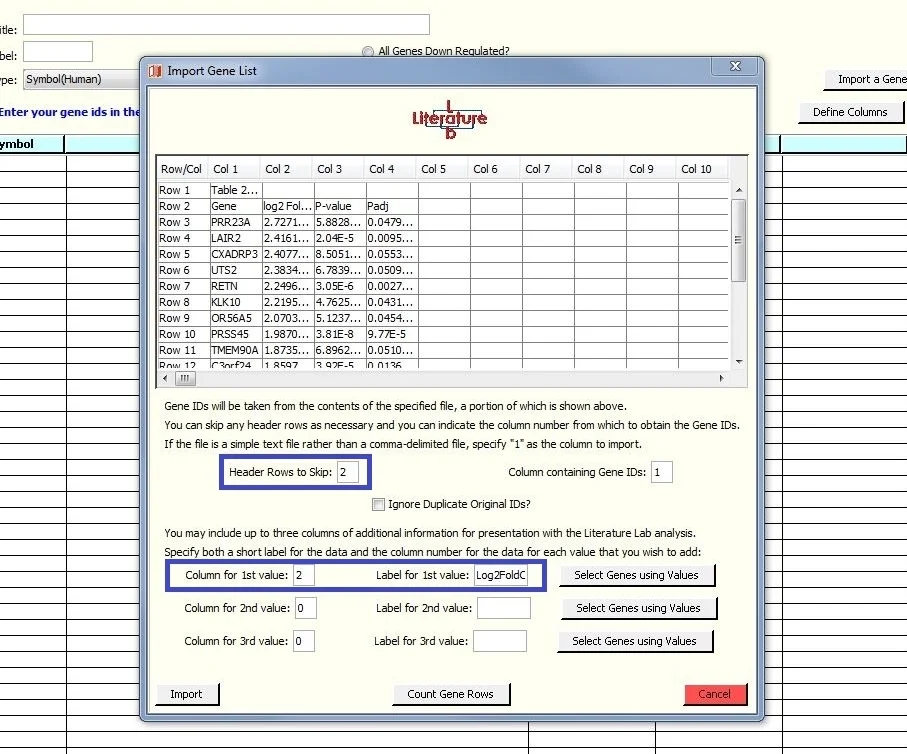
Specifying the additional data
We have specified that the data we want to import, in addition to the genes, is in column two, and the label is 'Log2FoldChg'.
We have also specified that the two header rows are not to be imported.
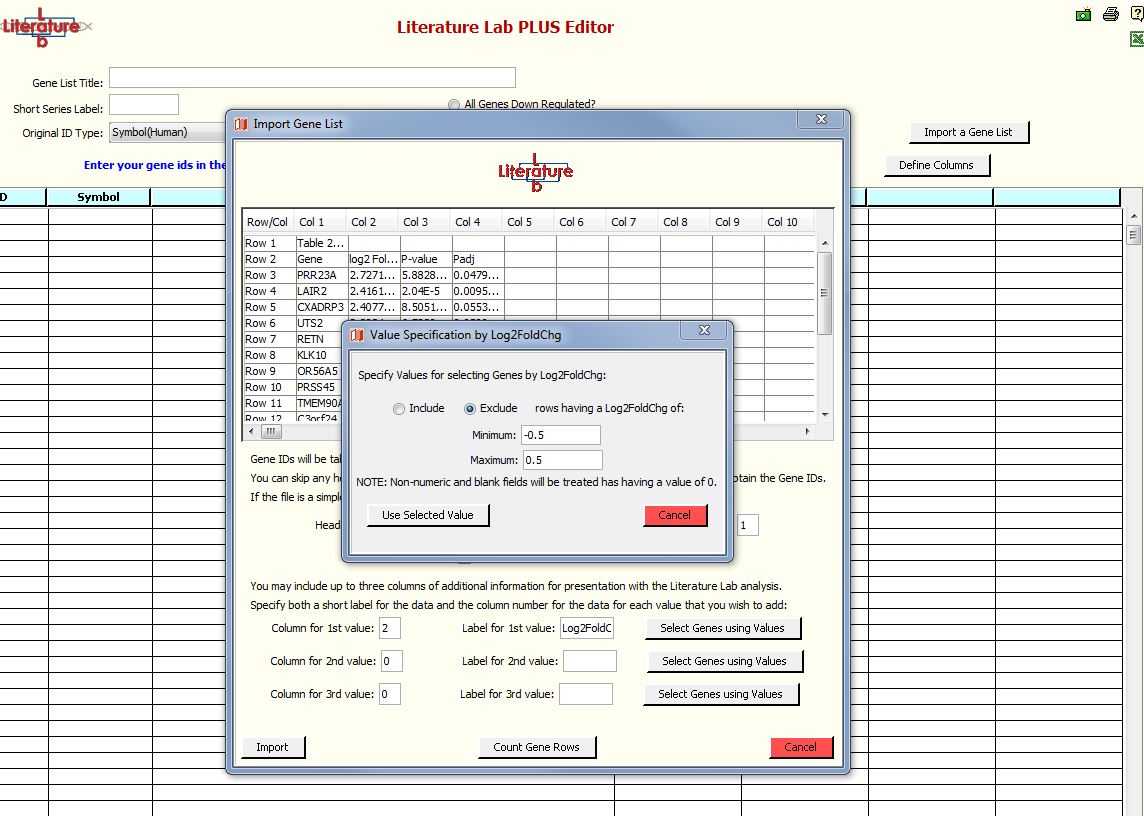
Specifying gene rows for removal
In this exercise, we will remove genes with fold change values between +0.5 and -0.5.
Click 'Select Genes Using Values'.
Select 'Exclude' and enter -0.5 and 0.5 into the Minimum and Maximum fields.
Click 'Use Selected Values'.
The Include and Exclude options enable other strategies for data selection. Selections can be made in all three columns or any single column.
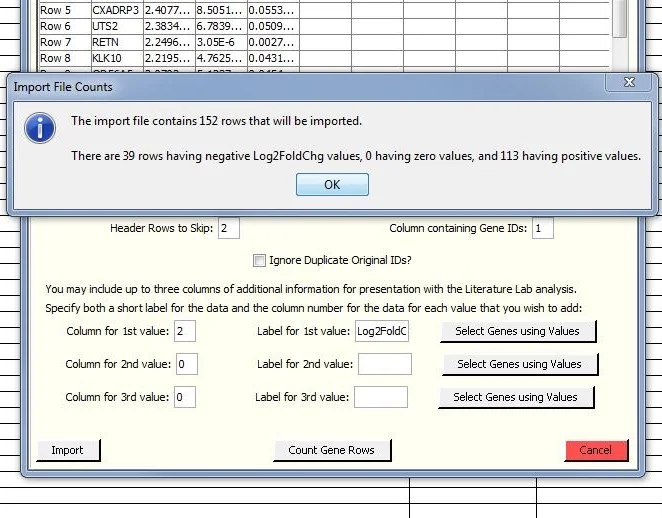
Reviewing our selection.
Click on 'Count gene Rows'.
This gives you a summary of the number of genes remaining with positive and negative fold change values. The "Select Genes Using Values" function can be used again if desired to evaluate the impact of other cutoff strategies.
When you are done specifying the fold change range, click 'Import' per instruction in Examples 1 & 2.
Validate the Gene list.
This view, ranked by fold change, shows that genes with fold change values between 0.5 and -0.5 have been removed.
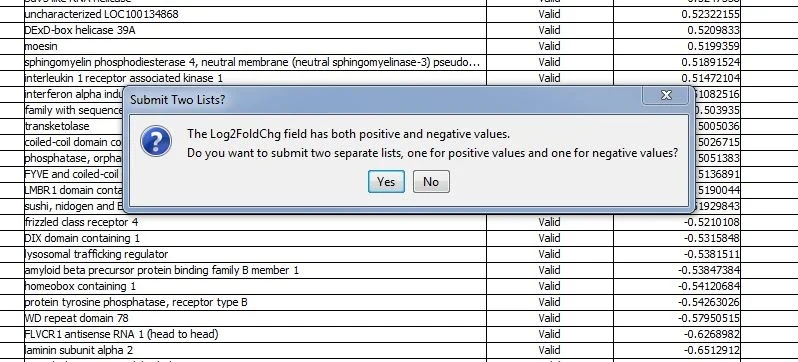
Next step: submit the data for Literature Lab PLUS analysis.
Click 'Yes'. Two gene lists will be submitted for analysis. '_Down' and '_Up' are appended to the file names.